Galaxy on HPC
Knowing that more and more often the data analysis request a lot of computational power, one of the challenges today is to be able to execute Galaxy on High Performance Computing clusters (HPC); several projects in different countries are running to solve this problem and to provide a Galaxy service for intensive computing.
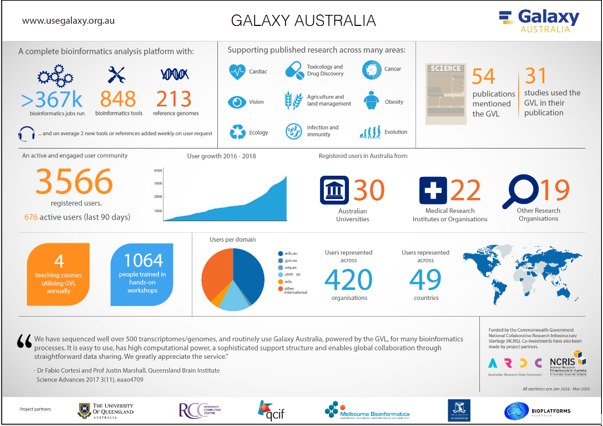
Galaxy Australia
- https://usegalaxy.org.au/
- https://static.sched.com/hosted_files/gcc2019/d5/S-1-2-GCC2019_GarethPrice.pdf
- Queensland Facility for Advanced Bioinformatics, Queensland Cyber Infrastructure Foundation, 306 Carmody Rd, St Lucia, Queensland 4067, Australia,
- Research Computing Centre, Level 5, Axon Building, The University of Queensland, St Lucia Queensland 4072, Australia
- Melbourne Bioinformatics, University of Melbourne, 187 Grattan Street, Victoria 3010, Australia
- Royal Botanic Gardens Private Bag 2000, Birdwood Avenue, South Yarra, Victoria 3141, Australia
- Centre for Comparative Genomics, Murdoch University, 90 South Street, Murdoch, Western Australia 6150, Australia
- Queensland Cyber Infrastructure Foundation, Level 5, Axon Building, The University of Queensland, St Lucia QLD 4072, Australia
Galaxy Australia (https://usegalaxy.org.au/) grew from three separate instances of Galaxy operated in Australia, in an effort to maximize resources (both human and computation) through reduced duplication and better operational synergies. Connected to Australian Data Repositories on National HPC, Galaxy Australia is used across many research areas: Cardiac, Vision, Ecology, Toxicology and Drug Discovery, Agriculture and land management, Infection and Immunity, Cancer, Obesity, Evolution.
AuBi
- https://mesocentre.uca.fr/projets-associes/plateforme-aubi/
- https://www.france-bioinformatique.fr/fr/plateformes/aubi
- https://static.sched.com/hosted_files/gcc2019/c0/GCC2019posterMesocentre.pdf
- Mésocentre Clermont Auvergne, Université Clermont Auvergne, 7 Avenue Blaise Pascal, 63 178 Aubière
- UMR454 MEDIS Microbiologie Environnement Digestif Santé, INRA Site de Theix 63 122 Saint-Genès-Champanelle, CBRV 26 Place Henri Dunant 63 001 Clermont-Ferrand
- Auvergne Bioinformatics platform, CBRV Faculté de Médecine & Pharmacie 28 Place Henri Dunant 63001 Clermont-Ferrand INRA Theix 63122 Saint-Genès-Champanelle
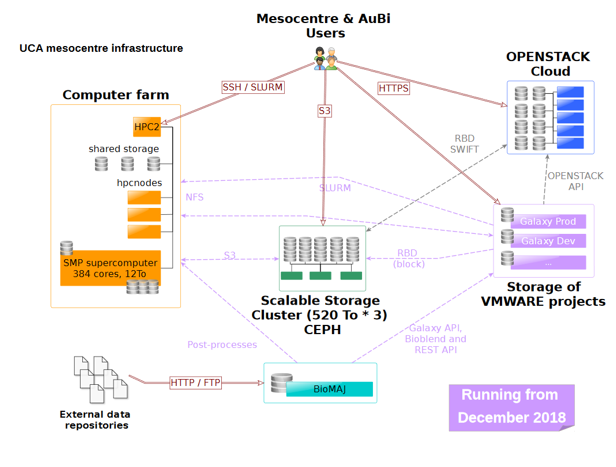
AuBi is the Auvergne Bioinformatics platform, hosted by Mesocentre (https://mesocentre.uca.fr). Mesocentre is delivering services in sciences data computing (HPC, VM, etc.) and short-term storage through a network of technology core facilities.
AuBi aims to share expertise and knowledge in large-scale data treatments and analysis by supplying a complete computing environment with hardware and software infrastructures for UCA (Clermont Auvergne University) research laboratories. From an informatics infrastructure point of view, a virtual machine, with an extensible disk on a scalable storage cluster (RBD / Ceph), runs the server. It is connected to the HPC facilities through a NFS server in order to allow Galaxy to benefit from the Mesocentre empowered computing and storage capabilities. Users authenticate through a shared LDAP directory between the Galaxy server and the cluster. In addition, BioMaJ framework (https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3293366/) was deployed for sharing databanks access to both Galaxy and cluster. In the future the access to the users from the IFB community via an eduGAIN connection will be open.
GenAP
https://static.sched.com/hosted_files/gcc2019/bf/Galaxy%20container%20in%20HPC.pdf
- Université de Sherbrooke, Sherbrooke, Quebec, Canada
- McGill University, Montreal, Quebec, Canada
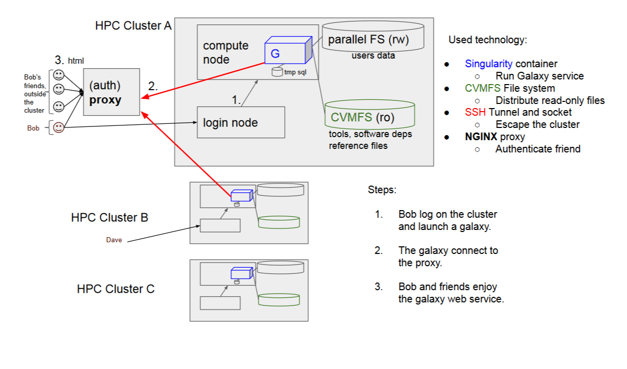
Containerizing Galaxy server as a Singularity application on HPC clusters for GenAP. The challenge is:
- Run Galaxy on every HPC (High Performance Computing) Cluster in Compute Canada.
- Run many, complete Galaxy services, on normal users accounts.
- Don’t alter the (old) cluster operating system, use the current setup.
- Don’t use “root”, don’t bother cluster system administrators.
- Be secure.
- Be nice to other actual “non-Galaxy” users.
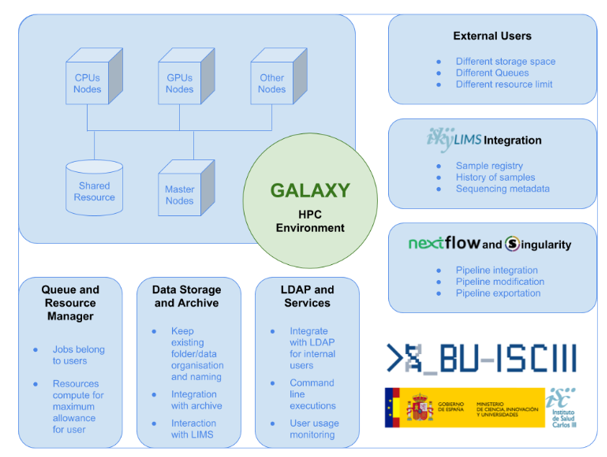
- Bioinformatics Unit(BU-ISCIII), Core facility, Health Institute Carlos III (ISCIII), Spain
- Genomics Unit, Core facility, Health Institute Carlos III (ISCIII), Spain
The Health Institute Carlos III (ISCIII) is the main Public Research Entity funding, managing and carrying out biomedical research in Spain. Galaxy is used in order to provide a unified and user friendly platform to execute standard genomics analysis, use custom tools and access the join pool of computational resources of research centers. Due to the complexity of the existing high performance computational (HPC) and data storage infrastructure, some parts of the standard Galaxy functionality will have to be extensively customized. In future the infrastructure and tools will be to external researchers, other smaller bioinformatics groups (like clinical bioinformatics units in hospitals) which face similar challenges and may lack the expertise or manpower to address them alone.